Food Safety
4 Advantages of Targeted DNA Sequencing over Automated Ribotyping - Test
Without subtyping, food manufacturers struggle to know if the Listeria they’ve detected today is the same Listeria they detected last week in a different location of the manufacturing facility.
To add to the struggle, understanding the microbial contamination of a facility is no easy task. Many companies currently rely on automated ribotyping to identify and track Listeria subtypes, but as we will discuss below, it’s not a complete solution.
Targeted DNA sequencing, alternatively, is a relatively new method–only available on the Clear Safety platform–that makes up for what automated ribotyping lacks, and offers several huge advantages to food companies.
The Difficulties with Legacy Subtyping Methods
For their environmental monitoring programs, many companies use the following workflow:
Listeria screening → Isolate Creation → Subtyping Method → Mapping Software
One of the weaknesses of this workflow is the subtyping component. First, legacy subtyping methods are labor-intensive. Since PFGE or ribotyping is the basis for pathogen characterization within many environmental monitoring programs, the turnaround time is painfully slow. Both methods are culture-dependent, and the generation of a pure isolate takes several days, which prolongs root cause analysis.
To learn more about how PFGE and ribotyping work, check out this resource, which compares PFGE, ribotyping, and targeted DNA sequencing.
Second, legacy subtyping methods are inaccurate, which, in turn, impacts operational efficiencies. To illustrate this point, let’s imagine that, when you receive your subtyping data, you find a specific subtype of Listeria in your packaging area. As you pore over the data, you realize that this subtype does not appear anywhere else in your facility.
To find the root cause, you could spend hours trying to determine how this unique subtype found its way to your packaging area. But what if this seemingly unique subtype is not a unique subtype, after all? What if it’s actually a subtype that you’re finding elsewhere in your facility but your current method mischaracterized the subtype? In other words, what if the problem has to do with the inaccuracy of your subtyping method? If that’s the case, your team may have spent undue time trying to understand a contamination incident that had a much simpler explanation.
The Advantages of Similarity Analysis™
Need a faster, more accurate solution? That’s where Similarity Analysis comes in.
Unlike whole-genome sequencing (WGS), which reads the entire genome, Similarity Analysis uses targeted sequencing to analyze specific regions of the genome. It then catalogs the genetic “fingerprint” of each Listeria encountered by the system and compares it to the fingerprints that were encountered previously.
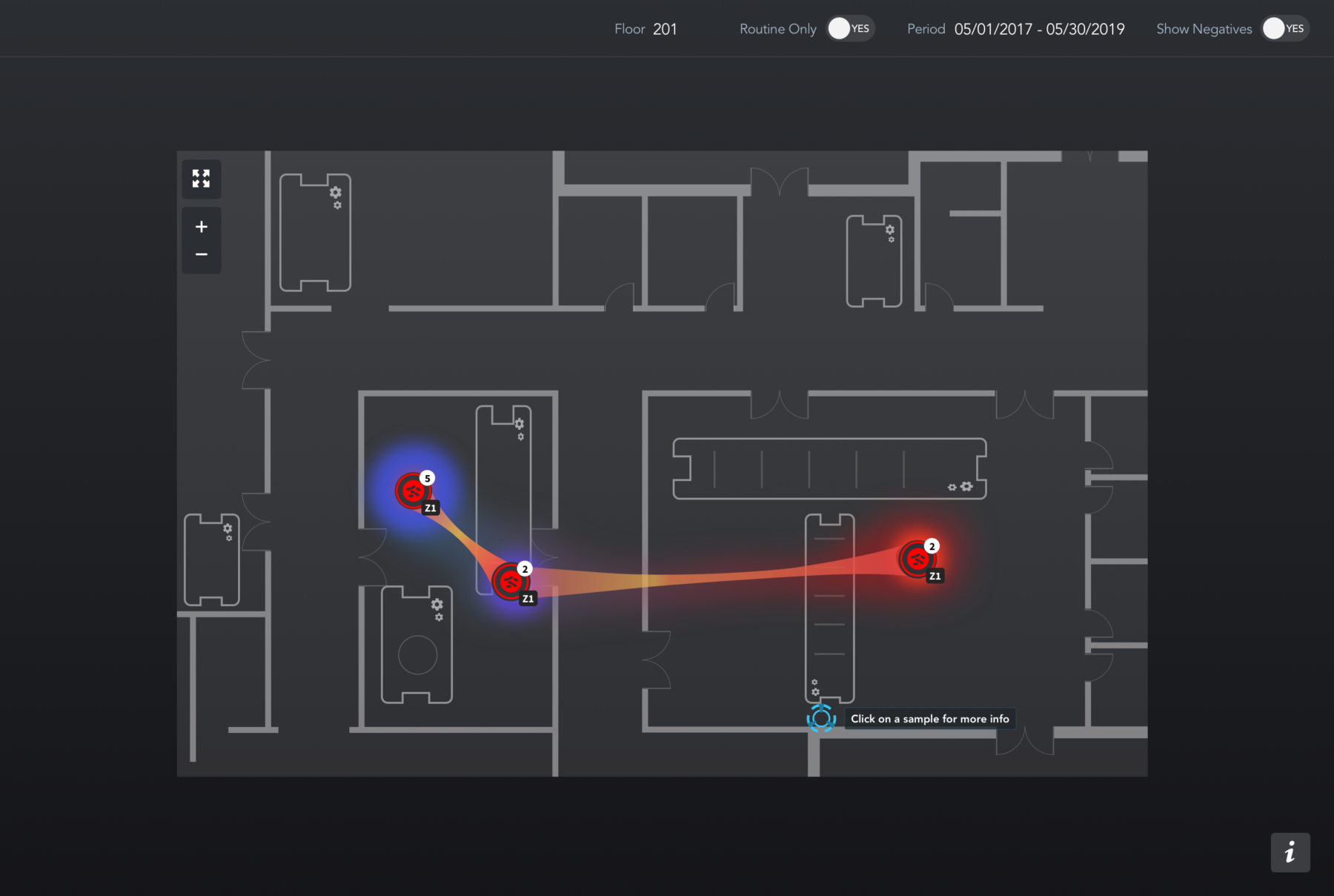
Samples containing Listeria with matching DNA fingerprints are grouped accordingly along with sample metadata, including the location, date, and time. This allows the user to track the flow of a particular type of Listeria through a plant over time.
1. No Isolates Required = Faster Turnaround Time
Currently, many food manufacturers rely on rapid typing methods requiring the creation of isolates. Isolation takes time. Clear Safety performs Similarity Analyses directly from enriched samples, improving your operational inefficiency and allowing you to identify the root cause of a contamination incident sooner.
2. Better Accuracy than Rapid Typing Methods
Whereas rapid typing methods look at a few distinct regions of the genome, Similarity Analysis looks at numerous regions spanning the genome. Analyzing more regions of the genome means that we have more information about the genome, and more information means that we can identify and characterize a microorganism more accurately.
3. Perform a Secondary Analysis
Up to 72 hours after an initial screening test, you can extract additional information from test data without re-processing the sample. For example, after an initial genus-level positive, you could choose to subtype your positive samples through Similarity Analysis.
4. Easy Integration
Currently, screening and subtyping results do not seamlessly flow into legacy mapping technology. Clear Safety changes all that, making it easier to visualize your results and providing you with new insights.
Ready to Get Started?
These are just four of the benefits of using Similarity Analysis for Listeria subtyping. If your business is still using legacy methods like PFGE or ribotyping, it’s time to ask yourself the following: Do you want faster turnaround times, improved efficiencies, better accuracy, and more flexibility?
If you do, contact us. We’d love to discuss how Clear Safety can help your food safety programs.